NORWAY SPRUCE FINE ROOTS
Main link
Secondary link
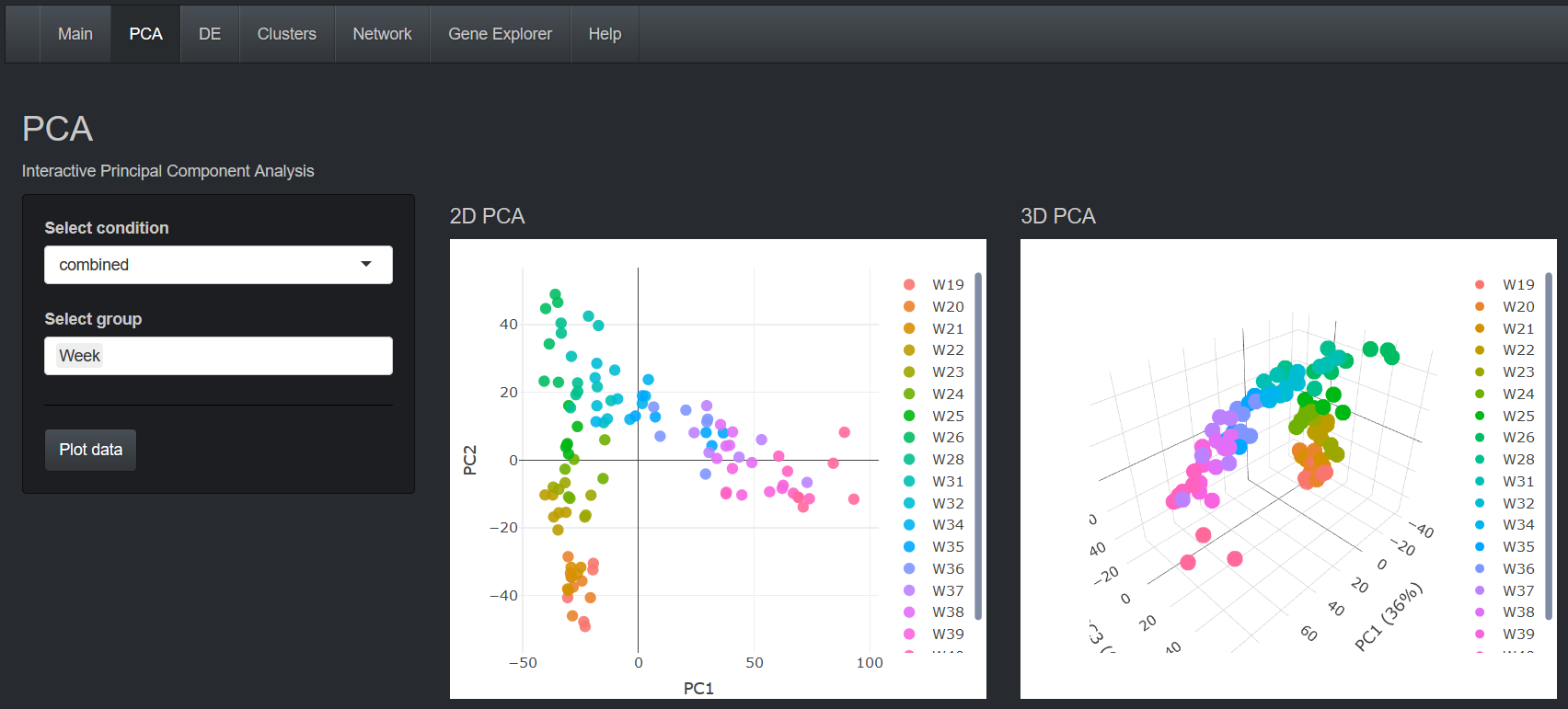
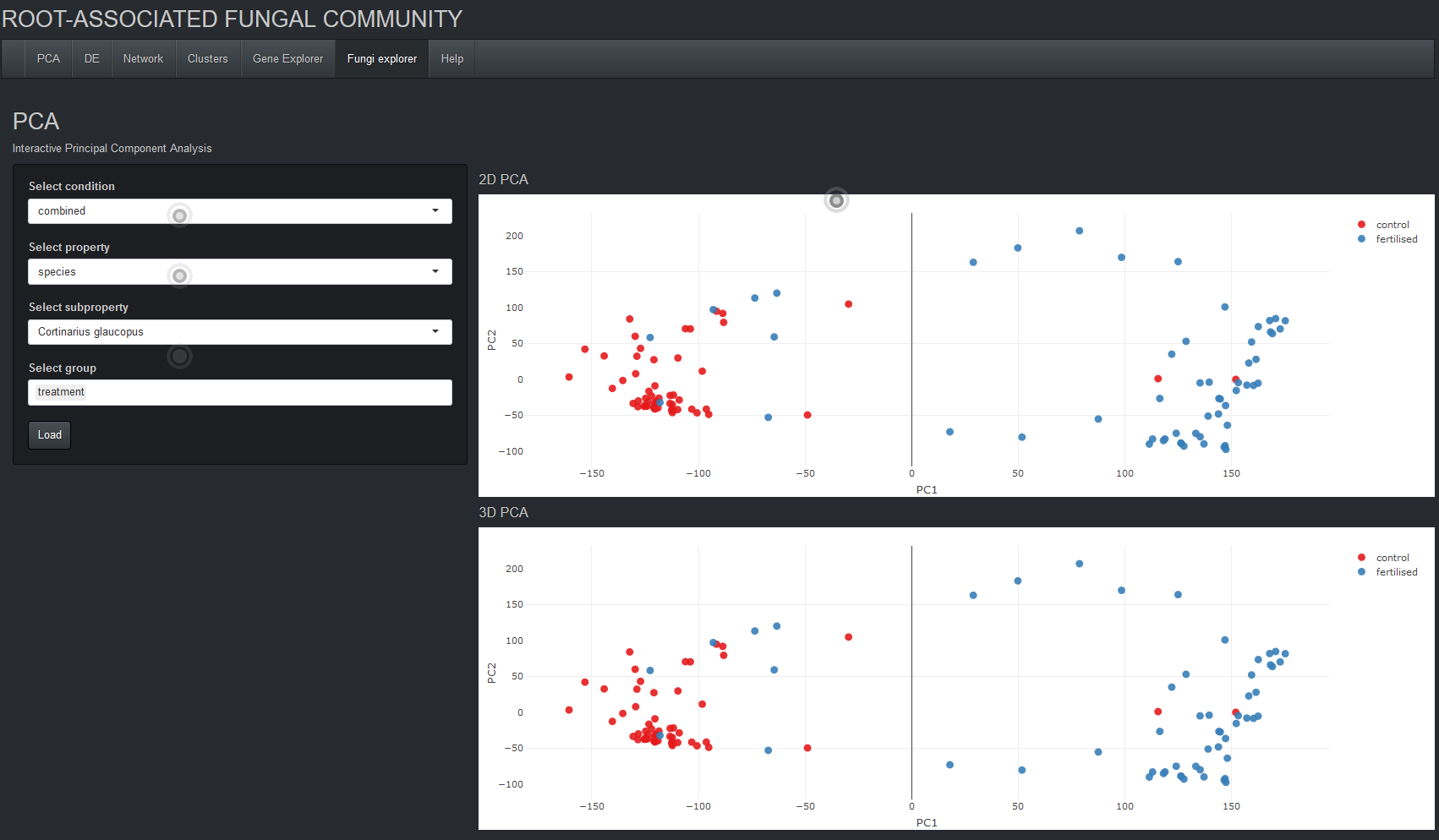
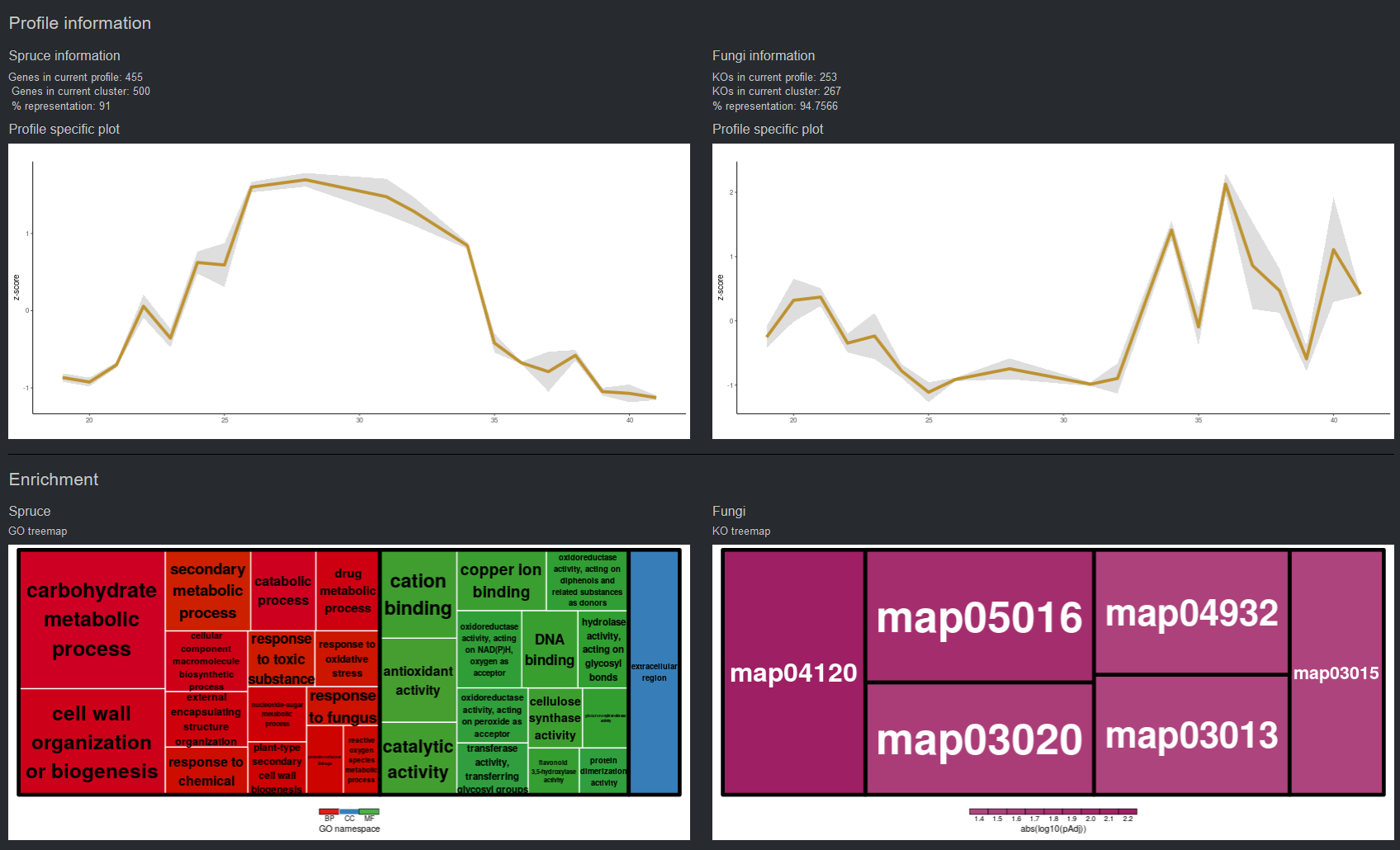
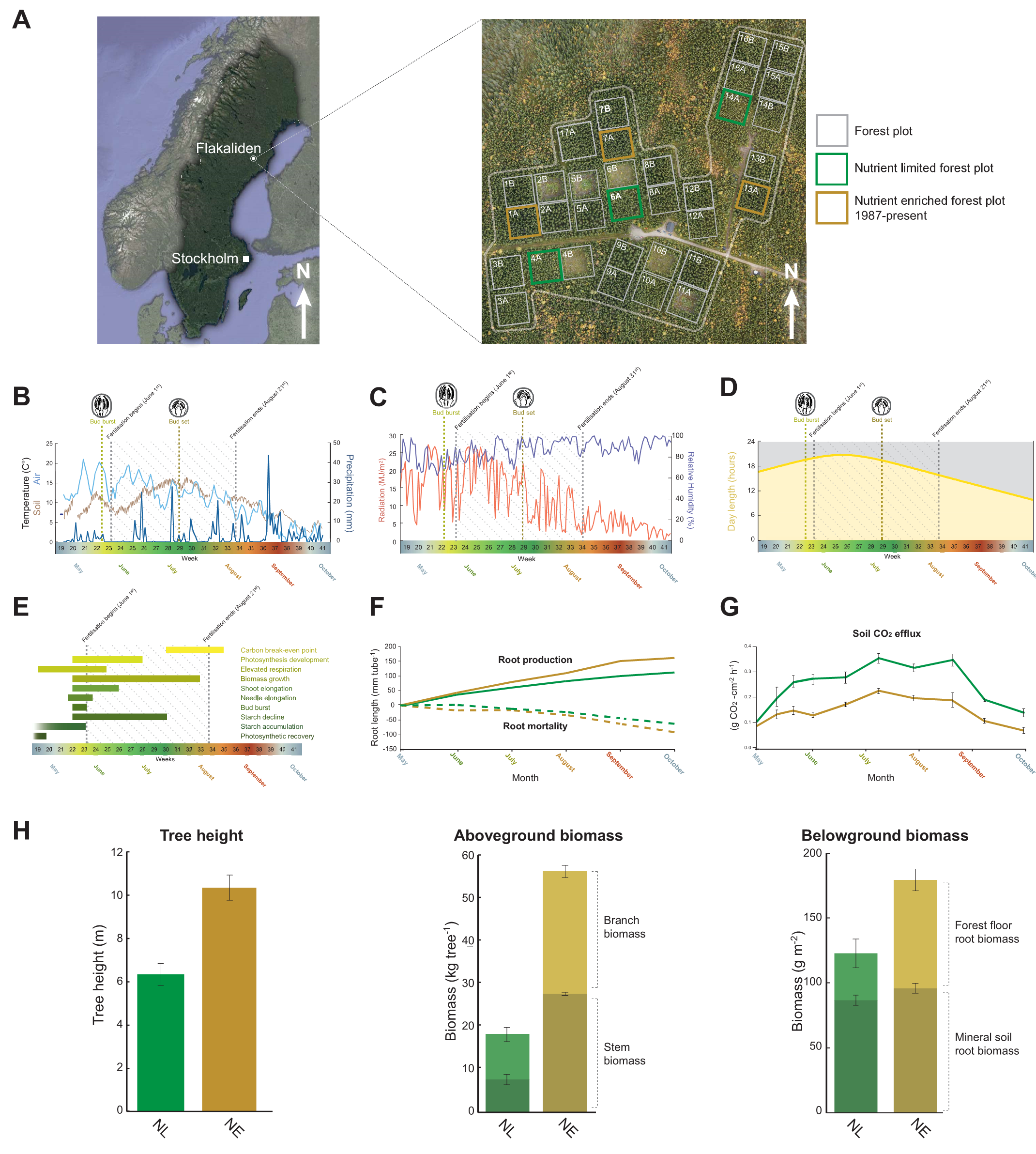
Temporally resolved transcriptomic analysis of Norway spruce (Picea abies) fine roots grown in nutrient-deficient
and nutrient-enriched forest plots in the boreal region of Northern Sweden.